Author: Jonathan French, ScD on March 12, 2019 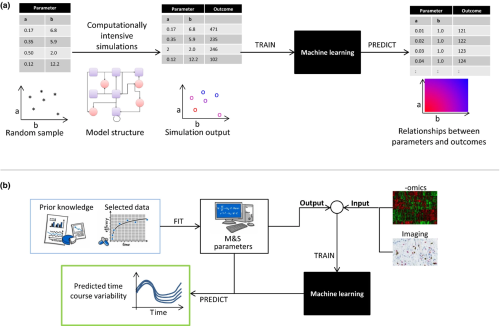
Machine learning (ML) and, specifically, deep learning (DL) algorithms are increasingly being used in the biological and health sciences to build predictive algorithms and to find associations between predictive features and outcomes. Examples of ML/DL applications are plentiful in areas as diverse as computational chemistry, observational data analysis of electronic health records, and the development of digital biomarkers.
Because of their success in other fields and access to higher computational power, incorporating ML/DL into the model-informed drug development process and clinical pharmacology decision-making is an area of active research. This activity was demonstrated by the number of abstracts and sessions at the recent ACOP9 meeting and other published applications, including Supervised Machine‐Learning Reveals That Old and Obese People Achieve Low Dapsone Concentrations by Hall et al.
Our modeling community is poised for a meaningful change in how modeling and simulation (M&S) is conducted to support clinical pharmacology decision-making as we incorporate ML/DL into the standard pharmacometrics and systems pharmacology toolkits.
In a recent perspective article, Hutchinson et al. discuss two illustrative examples in which the combination of ML and traditional M&S methods could yield a large improvement in model predictivity over either approach alone. One example focuses on machine learning for assisting in a global sensitivity analysis of complex quantitative systems pharmacology models. The second demonstrates how ML could be incorporated into the covariate search process in more traditional pharmacometric models.
Their paper, Models and Machines: How Deep Learning Will Take Clinical Pharmacology to the Next Level, provides two of what will be many examples in coming years. The authors and editors of CPT: Pharmacometrics & Systems Pharmacology hope that their paper stimulates discussion about this important topic.
Image by Hutchinson et al. CPT Pharmacometrics Syst. Pharmacol., doi: 10.1002/psp4.12377, is licensed under CC BY-NC-ND 4.0. ©2018 The authors.

The comment feature is locked by administrator.